Carnegie Mellon research suggests our view of COVID-19 is going to change with ‘digital surveillance’

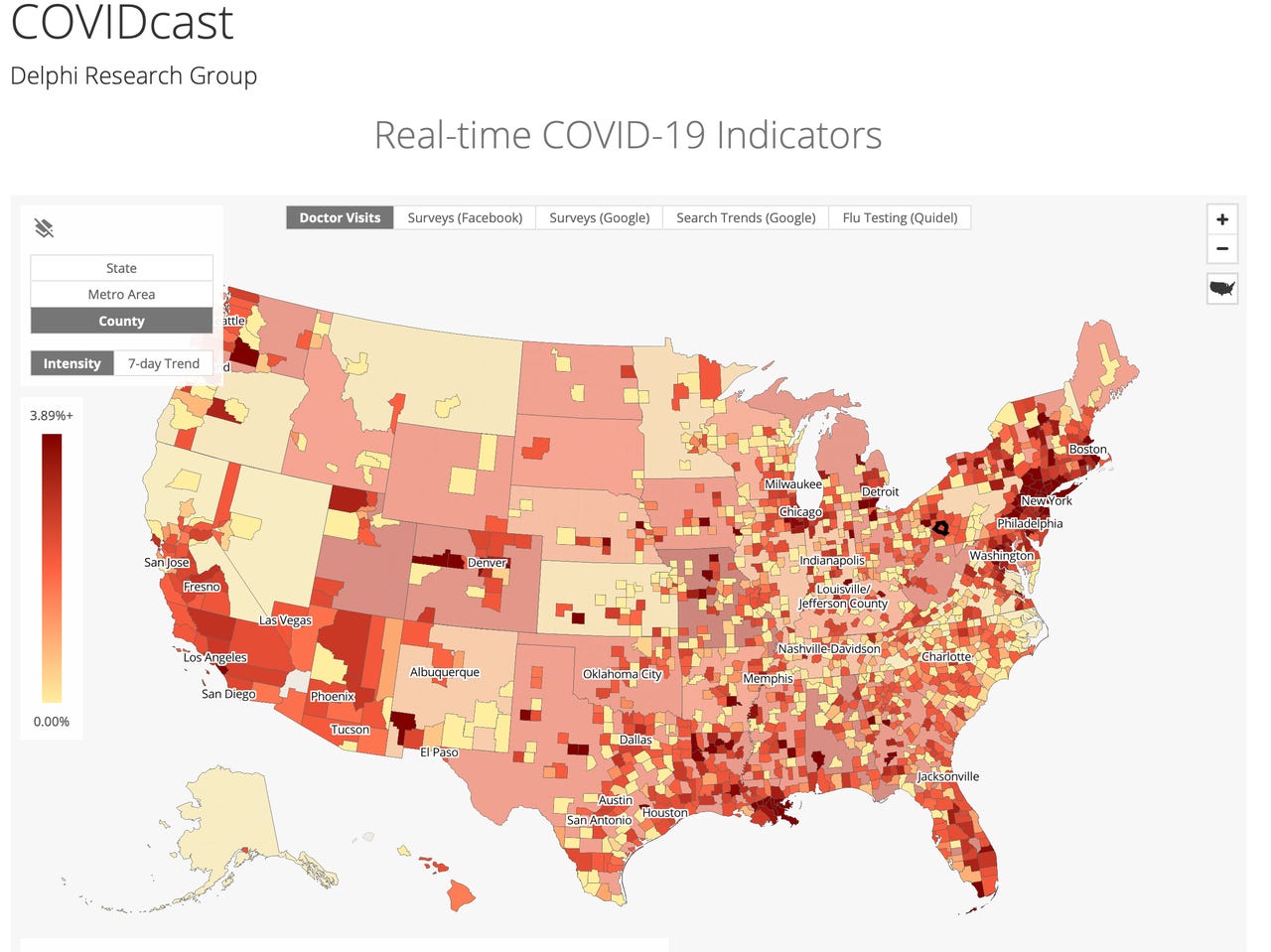
The COVIDcast Web site, assembled by the machine learning scholars of Carnegie Mellon's Delphi group, is combining multiple self-reported data points for the purpose of a "nowcast," a real-time picture of the disease that will be used to create new forecasts.
Correction: Quidel is providing tests for flu, not for COVID-19, as stated in a prior version of this article. The Delphi group expects to begin nowcasting in some weeks from now.
Doctors, researchers, and governments never have the full view of a disease outbreak at any given point in time. The full extent, indeed, may ultimately be unknowable, as scientists have pointed out.
And the extent of society's lack of knowledge is coming to light in the scrutiny of COVID-19 models.
New York governor Andrew Cuomo's daily briefings reveal the uncertainty in the multiple models of the disease. Popular media has pointed out where the models get it wrong, including "hidden" outbreaks spreading through American cities and vastly higher case counts than official tallies.
The problem is the way that COVID-19 has been estimated, and the forecasts that have been created, are a rather blunt instrument based on an epidemiological forecasting approach created 100 years ago.
That's about to change.
On Thursday, scientists at Carnegie Mellon University, one of two "centers of excellence" for research into influenza (the other being UMass at Amherst) unveiled a combination of five maps of symptoms reported all over the country by people who are feeling something that might be COVID-19, though it could be something else.
You can see the five maps at COVIDcast, the CMU website set up by the lab running the experiment, Delphi, lead by professors Roni Rosenfeld and Ryan Tibshirani, both of whom are from CMU's Department of machine learning. Tibshirani also has an appointment with CMU's Department of Statistics.
This combined symptom map at COVIDcast is an example of digital surveillance having real-time information by which to track the emergence of disease and to see its subtleties and nuances at the scale of localities. The sources include surveys filled out by Facebook users, users of Google Opinion Rewards, and Quidel Corp (a maker of medical tests, which records when people order a flu test, which can a possible indicator that a person is feeling symptoms akin to those of COVID-19).
The gathering of this data in real-time will eventually lead to what is called "nowcasting,"a practice that has evolved over the past decade as a way to get around the slow pace of which epidemiological data is accumulated. The COVID-19 team at Delphi has been perfecting nowcasting for eight years to predict seasonal influenza on behalf of the Centers for Disease Control.
Their approach for influenza relied on reported cases passed along from health care providers. This data, labeled ILINet, is a week old, meaning the patients that doctors see are not reported until the Friday of the following week.
The Delphi team developed nowcasting as a way to fuse real-time data such as Google search data about how many people search for symptoms of flu. Using a statistical approach, they amplify what the medical reports are saying by folding in what the real-time reports say.
The Delphi group is now building upon that expertise with the flu to create a new forecasting approach for COVID-19. Using the digital surveillance data of voluntary reporting, and the sensor fusion approach of combining multiple sources of data, they will start providing forecasts of the disease in the coming weeks based on the nowcast picture of things. Once they begin the forecasts, Delphi intend to add nowcasting of COVID-19.
The COVID-19 effort is not simply an extension of the influenza forecasting. It will require new statistical approaches because COVID-19 is not the same as seasonal influenza.
"What we're observing now is nothing like what we've observed with historical flu," Tibshirani remarked during a presentation at the COVID-19 and AI conference on April 1, sponsored by Stanford University's Institute for Human-Centered AΙ. "By definition of a pandemic, [COVID-19 is] nothing like what we've observed, period."
To analyze statistically the mass of surveillance data, the Delphi team has refined over many years a bunch of statistical and machine learning approaches. They include something called delta density, which incorporates approaches based on "Markov Processes," whereby a given state of affairs can be inferred from preceding states of affairs. Subsequent data can retrospectively be used to revise prior assumptions about states of affairs, in a way that continually refines a model of a disease.
There is a certain artistry to how the varying signals are combined, an ability to know how to handle and consider the data, as Tibshirani explained to ZDNet in an email.
"These signals are not measuring the same thing," Tibshirani wrote, "and they're not even drawn with respect to the same population (Facebook vs Google users)."
The Delphi group doesn't consider the signals from the surveys as "ground truth," a phrase in statistics that means objectively true. Rather, "it is their individual temporal trends that we're most interested in," wrote Tibshirani.
"For example, if in a given county we see both of these signals spike, then this can be a meaningful indicator of increased COVID-19 activity. Implicit in this is that their individual biases are constant over time, which is a reasonable assumption."
To understand the significance of the nowcasting effort, and the forecasts that will come from it, think about the limitations of the current models, including the models from Imperial College in London and Columbia University.
Primers
All these models are what are known as "mechanistic" models because they are based on a very general understanding of the mechanism by which all infectious diseases spread, the "transmission dynamics," as it sometimes referred to.
The mechanistic models in use today mostly derive from one mathematical approach, a model called a compartmental model. The most familiar form of the compartmental model is the so-called susceptible, infectious, recovered -- or SIR -- mathematical model. SIR was first introduced in 1927 by scientists William Ogilvy Kermack and Anderson Gray McKendrick.
The SIR model -- and the many variants that are used today, such as SEIR, which includes "exposed" people -- are sets of equations into which one plugs values for variables such as the number of currently infected people, in order to come up with a convincing curve for the progression of a disease. These models have come to dominate people's thinking about the spread of the disease. The Reich Lab at UMass, the other center for excellence, has brilliantly combined the models to show people the variance in their predictions.
These SIR-derived models have proven their worth over decades, but they have limitations. The most glaring limitation is the reliance of these models on data from official sources, such as the number of confirmed cases. The other big limitation is that the models are all fairly rigid, making similar assumptions about the mechanism by which disease spreads.
The statistical approach of the Delphi group's COVIDcast is likely to bring out the nuances of COVID-19. We already know that the disease strikes people in very different ways based on age and pre-existing chronic conditions such as obesity. We also know that there is the varying genetic make-up of the disease, with different strains mixing in the infected population, with different proteins giving the disease an antigenic signature that can vary over time.
As more and more data are incorporated as signals, more of the specificity of how COVID-19 spreads will come to light. Rather than looking at one single disease that spreads uniformly throughout the population, it's conceivable that doctors and governments may be looking at a complex of conditions that need to be tackled in different ways.
Epidemiology is going to learn about disease tracking from this pandemic, as a result of digital surveillance and nowcasting on the largest scale ever attempted. The practice of tracking and fighting disease may never be the same again.