Microservers: Helping unlock the body's secrets while beating data centre sprawl

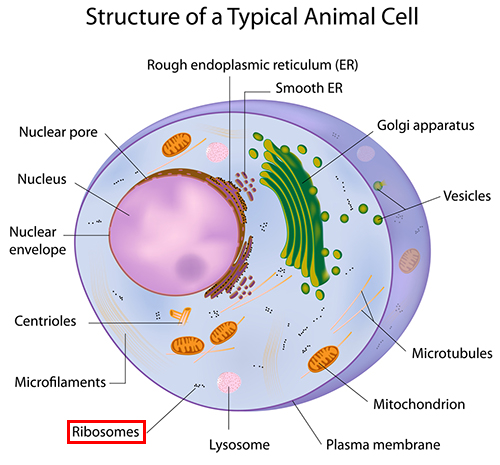
Inside every human cell are tiny protein factories churning out molecules that are crucial to our survival. Better understanding these biological replicators, known as ribosomes, could help to develop new pharmaceutical drugs. But getting a good look at ribosomes is tricky, because they measure just tens of nanometres across.
Ribosomes' tiny size means they have to be imaged using an electron microscope, which probes objects using a beam of electrons. Electron microscopes can resolve large amounts of detail, and are capable of magnifying objects up to 10,000,000 times compared to 2,000-times magnification for light-based apparatus.
Building a clear snapshot
Unfortunately, ribosomes can only withstand a relatively low dose of electrons before they are damaged by the intensity of the beam, which further complicates the imaging process. The modest dose they can endure only yields noisy, indistinct images.
"Even with this electron microscope you cannot really make a sharp image of them," said Andreas Hauser who worked as scientific computing expert at The Gene Center at the University of Munich in Germany, which has been studying ribosomes for 10 years.
Individually the images of ribosomes captured by the €4m cryo-electron microscope used in Munich are described as "more like a shadow".
To build a clear snapshot, the centre captures tens of thousands of photos of ribosomes as seen by the microscope. Each one of these digital images is fed into a computer, manipulated and combined to create a single, clearer composite picture of the ribosome's structure. Because the electron microscope captures the density, as well as the shape, of ribosomes, it's possible to build a 3D model.
"You get an image with say 80 ribosomes and you make 10,000 of those images, each with 80 ribosomes, so that's 800,000 particles," said Hauser. "You stamp out the part where the ribosomes are and you end up with one million very small images, like 300-by-300 pixels."
Each of these particles — images of individual ribosomes — is then aligned and combined into a single model.
The resulting model can provide a glimpse at fundamental processes of life, such as genetic instructions for building proteins passing into the ribosome.
"You can look inside this model and you can sometimes see RNA going in and out," said Hauser.
Beating datacentre sprawl
The Gene Center has been capturing images of ribosomes for 10 years and previously fed the images into groups of servers, providing up to 20 nodes per cluster.
As the equipment used to capture the images, known as micrographs, was improved however, so the computing equipment needed to keep pace. For example, higher-resolution imaging led the file size of micrographs to quadruple and the electron microscope was automated to capture snapshots around the clock.
The centre needed to upgrade, but it had little space to fit new racks and its maxed-out cooling meant it wasn't feasible to cram blade servers into existing racks.
"The server room was already quite full and we were thinking about how to fill the last rack unit," said Hauser.
"It was problematic because we thought if we go too dense our energy and cooling requirements would be too high. Our cooling was not made for more than 10-12KW per rack. If you go for these high-density blades you can end up with 20KW sometimes — and you need water-cooled racks, which we didn't have and hadn't planned for."
Instead the centre opted to buy two AMD SeaMicro SM10000 appliances, which provide dense clusters of server nodes with low-power consumption packed into a 10RU (Rack Unit) box.
"This was a good solution in terms of the space and the energy. You can easily fit three of the appliances in one rack with 10–12KW cooling."
"This was a good solution in terms of the space and the energy. You can easily fit three of the appliances in one rack with 10–12KW cooling."
— Andreas Hauser, The Gene Center, University of Munich
Hauser estimates using microserver clusters provided the same processing power as blade server clusters at between one half and one quarter the cooling cost.
Today the centre uses two SM10000 and two SM15000 appliances. These provide clusters of four-core Intel Xeon E3 and eight-core AMD Opteron processors, for a total of 1,256 processing cores.
Beyond cooling and space, the appliance also helped to meet the centre's networking need. Having sufficient bandwidth to shuttle images to server nodes is vital, with each image captured by the microscope weighing in at 4000-by-4000 pixels. "This terabyte of image data can cause a bottleneck getting it to the node," said Hauser. "With the SeaMicro nodes there are four cores and each node has an eight-gigabit link, so you get two gigabits per core. Usually if you have a 16-core server, even if you put in a relatively expensive 10-gigabit card, you don't even get one gigabit per core."
The throughput provided by the appliances is sufficient that the centre can avoid having to spend money on investing in Infiniband high-speed interconnects, added Hauser.
"The network can easily be more expensive than all of your computing hardware together. If you go for Infiniband, an Infiniband card is like €1,500 per node. Then you have to buy the switch and the cables — it's pretty harsh."
Each server node in a SeaMicro appliance is a PC server card that connects to the server backplane via PCI Express connection. The interconnect between the cards is dubbed the Freedom Fabric by SeaMicro, and provides 1.28Tbps of aggregate bandwidth. Each SM15000 appliance supports up to 16 10GbE or up to 64 1GbE external uplinks.
"It's like a PCI-Express network that allows for PCI-Express switching," said Hauser.
The cost of each SeaMicro appliance varies depending on the configuration, but Hauser estimates it ranges from €65,000 to €150,000, depending on the configuration. Compared to a blade setup Hauser estimates the upfront costs are the same, "but we save on space, on energy and on top of it we get the network for free".
Each server node in the appliances has 32GB of RAM and 50-150GB of disk space, which is used to store a local copy of images being processed. SeaMicro appliances supports up to 64 HDD or SSD drives.
The processing work needed to manipulate images captured by the microscope is suited to being handled by a cluster of less powerful servers, as it can be split into smaller tasks and processed in parallel.
"The good thing about our stuff is that it's easily parallelisable," said Hauser. "If you have one million particles and you divide them into batches of 1,000 it's often not such a big matter."
The SeaMicro clusters run Spider (System for Processing Image Data from Electron microscopy and Related fields) software on top of a modified version of Linux.
New drugs and discoveries
The ability to run more compute nodes 24/7 is helping researchers at The Gene Center, such as Roland Beckmann, combine larger numbers of individual images of ribosomes, referred to as particles, into more detailed and accurate models.
"Five years ago they maybe used 100,000 to 200,000 particles. Our institution was the first to use around one million particles," said Hauser. "With this increase in the number of particles we needed a lot more computational power, but the more particles you use the better the resolution is. We're now at four angstrom [a unit of measurement equivalent to one 10-billionth of a metre] and can distinguish between blobs of four atoms."
Building these rich models is important for understanding more about how ribosomes function — knowledge that could help develop pharmaceutical drugs with fewer side effects.
"We don't know how these small protein robots work in our cells and the 3D volume of a ribosome gives you an insight about their structure," said Hauser. "One application in about 10 years' time might be that a drug company develops medicines that work against the ribosomes of bacteria, but not the ribosomes in your cells.
"The more we know about what the ribosome looks like, the easier it gets to design drugs that only affect bacteria."